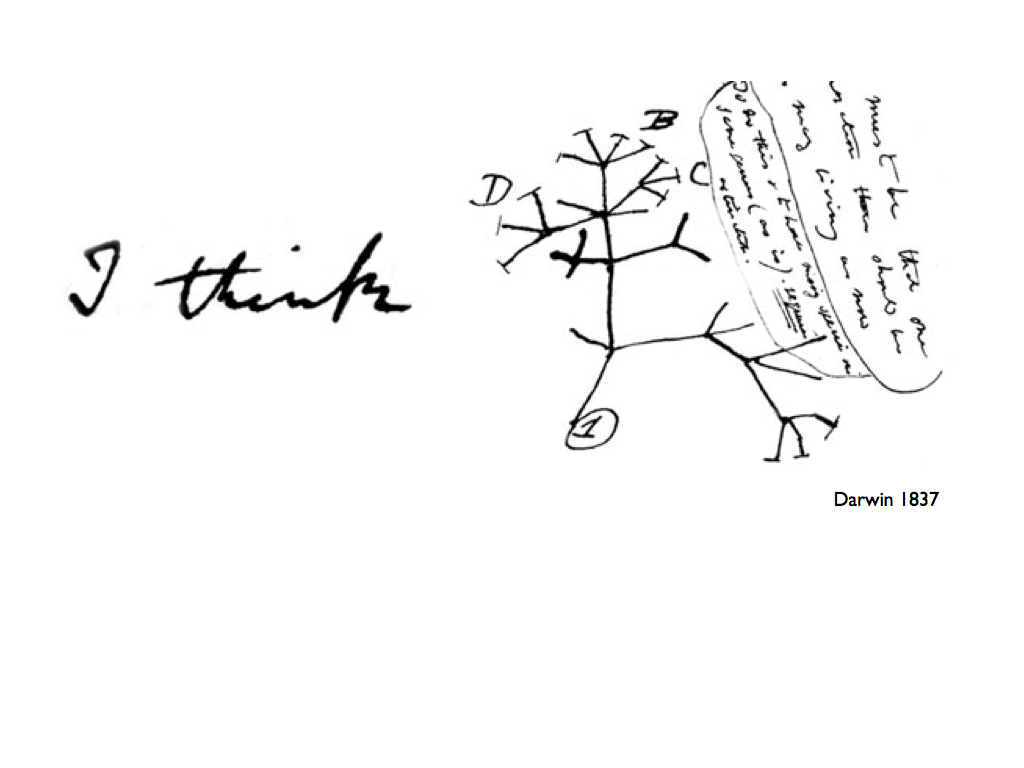
The first image we are used to associate to a phylogeny, is that of a tree (the tree of life, the genealogical tree of a family, the tree of languages..). What emerged in the last decade, and across different disciplines, is that this representation is too reductive in a wide number of cases. In biology, horizontal genes transfer is widely recognized to be a major mechanism of variation for many organisms. In linguistic, loans from a language to another play an important role in the evolution of a language. The identification of these phenomena induces a change in perspective with respect to the old idea of phylogenesis. In particular, in many cases, a clear hierarchy in the evolution lacks, so that phylogeny can no longer be represented as a tree and we have to consider some kind of network instead. This evidence uncovers the gap between the present and the needed analysis tools. While well established results are available for perfect phylogenies (i.e. evolutionary history that can be associated to a tree topology), when a deviation from a tree-like structure has to be considered very little is known, despite the efforts in this direction.
Our acivities can be ordered as follows.
2013 |
Taggi, Lorenzo; Colaiori, Francesca; Loreto, Vittorio; Tria, Francesca Dynamical correlations in the escape strategy of Influenza A virus (Journal Article) EUROPHYSICS LETTERS, 101 , 2013. (Abstract | BibTeX) @article{b,
title = {Dynamical correlations in the escape strategy of Influenza A virus},
author = {Lorenzo Taggi and Francesca Colaiori and Vittorio Loreto and Francesca Tria},
year = {2013},
date = {2013-01-01},
journal = {EUROPHYSICS LETTERS},
volume = {101},
abstract = {The evolutionary dynamics of human Influenza A virus presents a challenging theoretical problem. An extremely high mutation rate allows the virus to escape, at each epidemic season, the host immune protection elicited by previous infections. At the same time, at each given epidemic season a single quasi-species, that is a set of closely related strains, is observed. A non-trivial relation between the genetic (i.e., at the sequence level) and the antigenic (i.e., related to the host immune response) distances can shed light into this puzzle. In this paper we introduce a model in which, in accordance with experimental observations, a simple interaction rule based on spatial correlations among point mutations dynamically defines an immunity space in the space of sequences. We investigate the static and dynamic structure of this space and we discuss how it affects the dynamics of the virus-host interaction. Interestingly we observe a staggered time structure in the virus evolution as in the real Influenza evolutionary dynamics. © Copyright EPLA, 2013.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
The evolutionary dynamics of human Influenza A virus presents a challenging theoretical problem. An extremely high mutation rate allows the virus to escape, at each epidemic season, the host immune protection elicited by previous infections. At the same time, at each given epidemic season a single quasi-species, that is a set of closely related strains, is observed. A non-trivial relation between the genetic (i.e., at the sequence level) and the antigenic (i.e., related to the host immune response) distances can shed light into this puzzle. In this paper we introduce a model in which, in accordance with experimental observations, a simple interaction rule based on spatial correlations among point mutations dynamically defines an immunity space in the space of sequences. We investigate the static and dynamic structure of this space and we discuss how it affects the dynamics of the virus-host interaction. Interestingly we observe a staggered time structure in the virus evolution as in the real Influenza evolutionary dynamics. © Copyright EPLA, 2013. |
Tria, Francesca; Pompei, Simone; Loreto, Vittorio Dynamically correlated mutations drive human Influenza A evolution (Journal Article) SCIENTIFIC REPORTS, 3 , 2013. (Abstract | Links | BibTeX) @article{b,
title = {Dynamically correlated mutations drive human Influenza A evolution},
author = {Francesca Tria and Simone Pompei and Vittorio Loreto},
url = {http://www.nature.com/srep/2013/130919/srep02705/full/srep02705.html},
year = {2013},
date = {2013-01-01},
journal = {SCIENTIFIC REPORTS},
volume = {3},
publisher = {NATURE PUBLISHING GROUP},
abstract = {Human Influenza A virus undergoes recurrent changes in the hemagglutinin (HA) surface protein, primarily involved in the human antibody recognition. Relevant antigenic changes, enabling the virus to evade host immune response, have been recognized to occur in parallel to multiple mutations at antigenic sites in HA. Yet, the role of correlated mutations (epistasis) in driving the molecular evolution of the virus still represents a challenging puzzle. Further, though circulation at a global geographic level is key for the survival of Influenza A, its role in shaping the viral phylodynamics remains largely unexplored. Here we show, through a sequence based epidemiological model, that epistatic effects between amino acids substitutions, coupled with a reservoir that mimics worldwide circulating viruses, are key determinants that drive human Influenza A evolution. Our approach explains all the up-to-date observations characterizing the evolution of H3N2 subtype, including phylogenetic properties, nucleotide fixation patterns, and composition of antigenic clusters.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
Human Influenza A virus undergoes recurrent changes in the hemagglutinin (HA) surface protein, primarily involved in the human antibody recognition. Relevant antigenic changes, enabling the virus to evade host immune response, have been recognized to occur in parallel to multiple mutations at antigenic sites in HA. Yet, the role of correlated mutations (epistasis) in driving the molecular evolution of the virus still represents a challenging puzzle. Further, though circulation at a global geographic level is key for the survival of Influenza A, its role in shaping the viral phylodynamics remains largely unexplored. Here we show, through a sequence based epidemiological model, that epistatic effects between amino acids substitutions, coupled with a reservoir that mimics worldwide circulating viruses, are key determinants that drive human Influenza A evolution. Our approach explains all the up-to-date observations characterizing the evolution of H3N2 subtype, including phylogenetic properties, nucleotide fixation patterns, and composition of antigenic clusters. |
2012 |
Pompei, Simone; Loreto, Vittorio; Tria, Francesca Phylogenetic Properties of RNA Viruses (Journal Article) PLOS ONE, 7 , pp. e44849-1–e44849-10, 2012. (Abstract | Links | BibTeX) @article{b,
title = {Phylogenetic Properties of RNA Viruses},
author = {Simone Pompei and Vittorio Loreto and Francesca Tria},
url = {http://www.plosone.org/article/info%3Adoi%2F10.1371%2Fjournal.pone.0044849},
year = {2012},
date = {2012-01-01},
journal = {PLOS ONE},
volume = {7},
pages = {e44849-1--e44849-10},
abstract = {A new word, phylodynamics, was coined to emphasize the interconnection between phylogenetic properties, as observed for instance in a phylogenetic tree, and the epidemic dynamics of viruses, where selection, mediated by the host immune response, and transmission play a crucial role. The challenges faced when investigating the evolution of RNA viruses call for a virtuous loop of data collection, data analysis and modeling. This already resulted both in the collection of massive sequences databases and in the formulation of hypotheses on the main mechanisms driving qualitative differences observed in the (reconstructed) evolutionary patterns of different RNA viruses. Qualitatively, it has been observed that selection driven by the host immune response induces an uneven survival ability among co-existing strains. As a consequence, the imbalance level of the phylogenetic tree is manifestly more pronounced if compared to the case when the interaction with the host immune system does not play a central role in the evolutive dynamics. While many imbalance metrics have been introduced, reliable methods to discriminate in a quantitative way different level of imbalance are still lacking. In our work, we reconstruct and analyze the phylogenetic trees of six RNA viruses, with a special emphasis on the human Influenza A virus, due to its relevance for vaccine preparation as well as for the theoretical challenges it poses due to its peculiar evolutionary dynamics. We focus in particular on topological properties. We point out the limitation featured by standard imbalance metrics, and we introduce a new methodology with which we assign the correct imbalance level of the phylogenetic trees, in agreement with the phylodynamics of the viruses. Our thorough quantitative analysis allows for a deeper understanding of the evolutionary dynamics of the considered RNA viruses, which is crucial in order to provide a valuable framework for a quantitative assessment of theoretical predictions.},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
A new word, phylodynamics, was coined to emphasize the interconnection between phylogenetic properties, as observed for instance in a phylogenetic tree, and the epidemic dynamics of viruses, where selection, mediated by the host immune response, and transmission play a crucial role. The challenges faced when investigating the evolution of RNA viruses call for a virtuous loop of data collection, data analysis and modeling. This already resulted both in the collection of massive sequences databases and in the formulation of hypotheses on the main mechanisms driving qualitative differences observed in the (reconstructed) evolutionary patterns of different RNA viruses. Qualitatively, it has been observed that selection driven by the host immune response induces an uneven survival ability among co-existing strains. As a consequence, the imbalance level of the phylogenetic tree is manifestly more pronounced if compared to the case when the interaction with the host immune system does not play a central role in the evolutive dynamics. While many imbalance metrics have been introduced, reliable methods to discriminate in a quantitative way different level of imbalance are still lacking. In our work, we reconstruct and analyze the phylogenetic trees of six RNA viruses, with a special emphasis on the human Influenza A virus, due to its relevance for vaccine preparation as well as for the theoretical challenges it poses due to its peculiar evolutionary dynamics. We focus in particular on topological properties. We point out the limitation featured by standard imbalance metrics, and we introduce a new methodology with which we assign the correct imbalance level of the phylogenetic trees, in agreement with the phylodynamics of the viruses. Our thorough quantitative analysis allows for a deeper understanding of the evolutionary dynamics of the considered RNA viruses, which is crucial in order to provide a valuable framework for a quantitative assessment of theoretical predictions. |
2011 |
Pompei, Simone; Tria, Francesca; Loreto, Vittorio On the accuracy of language trees (Journal Article) PLOS ONE, 6(6) , 2011. (Links | BibTeX) @article{b,
title = {On the accuracy of language trees},
author = {Simone Pompei and Francesca Tria and Vittorio Loreto},
url = {http://www.scopus.com/inward/record.url?eid=2-s2.0-79958041006&partnerID=65&md5=22335465a4b96cbddf4bbc9cefd7b7a8
http://samarcanda.phys.uniroma1.it/vittorioloreto/PAPERS/2011/POMPEI_PLOS_ONE_2011.pdf},
year = {2011},
date = {2011-01-01},
journal = {PLOS ONE},
volume = {6(6)},
publisher = {San Francisco, CA : Public Library of Science},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
2010 |
Tria, Francesca; Caglioti, Emanuele; Loreto, Vittorio; Pagnani, Andrea A Stochastic Local Search Algorithm for Distance-Based Phylogeny Reconstruction (Journal Article) MOLECULAR BIOLOGY AND EVOLUTION, 27 (11) , pp. 2587–2595, 2010. (Links | BibTeX) @article{b,
title = {A Stochastic Local Search Algorithm for Distance-Based Phylogeny Reconstruction},
author = {Francesca Tria and Emanuele Caglioti and Vittorio Loreto and Andrea Pagnani},
url = {http://www.scopus.com/inward/record.url?eid=2-s2.0-77958142872&partnerID=65&md5=8bb15034042b6d808b9afb8819bb44d6
http://samarcanda.phys.uniroma1.it/vittorioloreto/PAPERS/2010/Tria_Mol_Biol_Evol_2010.pdf},
year = {2010},
date = {2010-01-01},
journal = {MOLECULAR BIOLOGY AND EVOLUTION},
volume = {27 (11)},
pages = {2587--2595},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Tria, Francesca; Caglioti, Emanuele; Loreto, Vittorio; Pagnani, Andrea A stochastic local search approach to language tree reconstruction (Journal Article) 27 , pp. 341–358, 2010. (Links | BibTeX) @article{b,
title = {A stochastic local search approach to language tree reconstruction},
author = {Francesca Tria and Emanuele Caglioti and Vittorio Loreto and Andrea Pagnani},
url = {http://www.scopus.com/inward/record.url?eid=2-s2.0-77958505543&partnerID=65&md5=2a15ef158e89c0e12ba7d0efb7201de5, http://gateway.webofknowledge.com/gateway/Gateway.cgi?GWVersion=2&SrcApp=PARTNER_APP&SrcAuth=LinksAMR&KeyUT=000283796800009&DestLinkType=FullRecord&DestApp=ALL_WOS&UsrCustomerID=0c7ff228ccbaaa74236f48834a34396a},
year = {2010},
date = {2010-01-01},
volume = {27},
pages = {341--358},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Caglioti, Emanuele; Loreto, Vittorio; Pompei, Simone; Tria, Francesca Distance-based Phylogenetic algorithms: new insights and applications (Journal Article) MATHEMATICAL MODELS AND METHODS IN APPLIED SCIENCES, 20 , pp. 1511–1532, 2010. (Links | BibTeX) @article{b,
title = {Distance-based Phylogenetic algorithms: new insights and applications},
author = {Emanuele Caglioti and Vittorio Loreto and Simone Pompei and Francesca Tria},
url = {http://www.scopus.com/inward/record.url?eid=2-s2.0-77957171910&partnerID=65&md5=41f74dd5dff0e500c1061a8d02fcd6c8
http://samarcanda.phys.uniroma1.it/vittorioloreto/PAPERS/2010/POMPEI_M3A_2010.pdf},
year = {2010},
date = {2010-01-01},
journal = {MATHEMATICAL MODELS AND METHODS IN APPLIED SCIENCES},
volume = {20},
pages = {1511--1532},
publisher = {World Scientific Publishing Company:PO Box 128, Farrer Road, Singapore 912805 Singapore:011 65 6 4665775, EMAIL: journal@wspc.com.sg, INTERNET: http://www.wspc.com.sg, http://www.worldscinet.com, Fax: 011 65 6 4677667},
keywords = {},
pubstate = {published},
tppubtype = {article}
}
|
Tria, Francesca; Caglioti, Emanuele; Loreto, Vittorio; Pompei, Simone A fast noise reduction driven distance-based phylogenetic algorithm (Inproceeding) Hamid R. Arabnia Quoc-Nam Tran, Rui Chang Matthew He Andy Marsh Ashu Solo Jack Yang (Eds.) (Ed.): Proceedings of BIOCOMP 2010 (2010), pp. 375–380, CSREA Press 2010, 2010. (Links | BibTeX) @inproceedings{b,
title = {A fast noise reduction driven distance-based phylogenetic algorithm},
author = {Francesca Tria and Emanuele Caglioti and Vittorio Loreto and Simone Pompei},
editor = {Hamid R. Arabnia, Quoc-Nam Tran, Rui Chang, Matthew He, Andy Marsh, Ashu M. G. Solo, Jack Y. Yang (Eds.)},
url = {http://dblp.uni-trier.de},
year = {2010},
date = {2010-01-01},
booktitle = {Proceedings of BIOCOMP 2010 (2010)},
pages = {375--380},
publisher = {CSREA Press 2010},
keywords = {},
pubstate = {published},
tppubtype = {inproceedings}
}
|